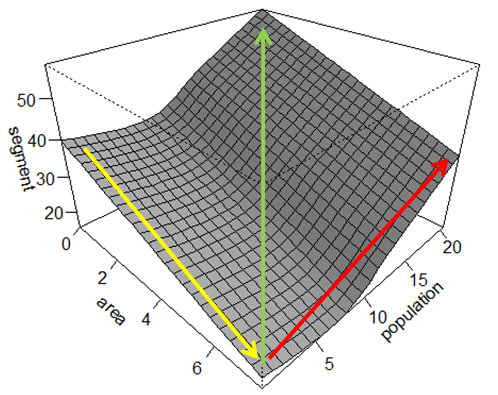
Right, I already referred to Atkinson’s paper in a previous post, and much of the work he’s presented is essentially part of a potential PhD project I’m hoping to do. Much of this stems back to last summer, where I mentioned how the phoneme inventory size correlates with certain demographic features, such as population size and population density. Using the the UPSID data I generated a generalised additive model to demonstrate how area and population size interact in determining the phoneme inventory size:
Interestingly, Atkinson seems to derive much of his thinking, at least in his choice of demographic variables, from work into the transmission of cultural artefacts (see here and here). For me, there are clear uses for these demographic models in testing hypotheses for linguistic transmission and change, as I see language as a cultural product. It appears Atkinson reached the same conclusion. Where we depart, however, is in our overall explanations of the data. My major problem with the claim is theoretical: he hasn’t ruled out other historical-evolutionary explanations for these patterns.
Before we get into the bulk of my criticism, I’ll provide a very brief overview of the paper.